04/23/2015
Objective: Run PCR product out on a gel to verify that the Luciferase cDNA was succesfully replicated
Results: Controls in lane 4 and 5 show contamination so will redo pcr with fresh ingredients (primers, h20, apex red etc)
PCR gel
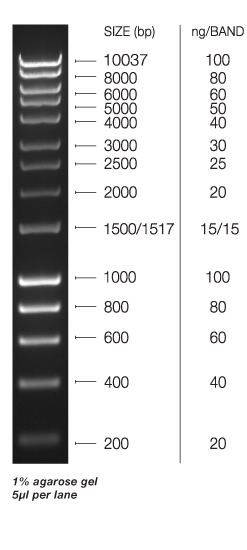
Ladder: o'gene ruler 100bp #sm1143 lot 00189314 0.1 ug/ul
Cells
1 5 ul Ladder
2 23ul cDNA 1
3 23ul CDNA 2
4 23ul H20 1 (control)
5 23ul H20 2 (control)
Thursday, April 23, 2015
Wednesday, April 22, 2015
04/22/2015
Objective: Ran pcr with template cDNA created from 4 rna sea pen rna samples on 04/15/2015 with Sam at the Roberts lab
used following temp. settings for the thermalcycler
Conventional PCR (2x Apex Red) [Cost per rxn ~ $0.52]
Single reaction (25uL) set up is listed below. Be sure to make a master mix volume that will accommodate all of your samples, two water (no template controls; NTC) samples, plus an extra 10% to accommodate pipetting errors. Distribute appropriate amount of master mix (volume of master mix + template = 25uL) to PCR tubes or PCR plate. Make sure all tubes/caps are tightly closed. Put in thermalcycler.
Reaction_Components
Volume
Final Concentration
2x Apex Red
12.5
1x
Forward Primer (10uM)
0.5
0.2uM
Reverse Primer (10uM)
0.5
0.2uM
Template
Up to 5uL
H2O (PCR grade)
variable
Use to bring reaction volume up to 25uL
Typical cycling paramaters (ask for help on using the thermal cycler):
95C - 10mins
40 cyles of:
95C - 15s
55C - 15s
72C - 1 mins (dependent on amplicon size; ~1000kb/min)
modified quantity measurements
Objective: Ran pcr with template cDNA created from 4 rna sea pen rna samples on 04/15/2015 with Sam at the Roberts lab
used following temp. settings for the thermalcycler
Conventional PCR (2x Apex Red) [Cost per rxn ~ $0.52]
Single reaction (25uL) set up is listed below. Be sure to make a master mix volume that will accommodate all of your samples, two water (no template controls; NTC) samples, plus an extra 10% to accommodate pipetting errors. Distribute appropriate amount of master mix (volume of master mix + template = 25uL) to PCR tubes or PCR plate. Make sure all tubes/caps are tightly closed. Put in thermalcycler.
Reaction_Components
Volume
Final Concentration
2x Apex Red
12.5
1x
Forward Primer (10uM)
0.5
0.2uM
Reverse Primer (10uM)
0.5
0.2uM
Template
Up to 5uL
H2O (PCR grade)
variable
Use to bring reaction volume up to 25uL
Typical cycling paramaters (ask for help on using the thermal cycler):
95C - 10mins
40 cyles of:
95C - 15s
55C - 15s
72C - 1 mins (dependent on amplicon size; ~1000kb/min)
modified quantity measurements
Results: Sucessfully completed PCR will run out on gel to veirfy replication of cDNA.
Wednesday, April 15, 2015
04/15/2015
Objective: Convert RNA to cDNA with Sam at the Friedman lab used following reverse transcription protocol
Methods:
Reverse Transcription (Promega M-MLV: Cat#M1701; ) [Cost per sample ~$1.50]
A single reaction volume = 25uL. The volume of RNA, primer(s) and M-MLV RT used are variable and will be specific to your current experiment. The directions below apply to a reaction using 1ug of total RNA. You may need to make changes to accommodate your own conditions.
1. Use as much RNA as possible (up to 1ug); max volume of RNA = 17.75uL. Generally, identify the RNA sample with the lowest concentration and multiply by 17.75uL. Use this quantity (ug) of RNA for each and every sample.
2. Transfer calculated volume(s) of RNA to 0.5mL snap cap tubes or PCR plate. Adjust volumes of individual samples to 17.75uL with H2O.
3. Add appropriate amount of primer to sample. Use 0.25ug primer per 1ug of RNA in sample (= 0.5uL of Promega oligo dT Cat#C1101 in this example). Total volume (RNA + primers) should equal 18.25uL.
4. Heat samples at 70C for 5 min in thermocycler.
5. Place samples on ice IMMEDIATELY.
6. Make Master Mix:
PER RXN
5 uL 5x Buffer (M-MLV RT Buffer)
1.25 uL 10mM dNTPs (Promega Cat#U1511)
0.5 uL M-MLV RT per ug of RNA
Used 4uL of RNA (
1ul of rna from each of the original sea pen sample added to a single sample combining rna sp1-1 sp1-2 sp2-1 sp2-2)
7. Mix well.
8. Add 6.75uL of master mix to each reaction.
9. Mix well, but do not vortex.
10.Spot spin.
11.Incubate @ 42C for 1hr in thermalcycler for oligo dT primers OR @ 37C for random primers.
12.Heat inactivate @ 95C for 3 min.
13.Spot spin.
14.Store @ -20C.
Sucessfully converted RNA to cDNA next step is to replicate using PCR and verify on a gel.
Objective: Convert RNA to cDNA with Sam at the Friedman lab used following reverse transcription protocol
Methods:
Reverse Transcription (Promega M-MLV: Cat#M1701; ) [Cost per sample ~$1.50]
A single reaction volume = 25uL. The volume of RNA, primer(s) and M-MLV RT used are variable and will be specific to your current experiment. The directions below apply to a reaction using 1ug of total RNA. You may need to make changes to accommodate your own conditions.
1. Use as much RNA as possible (up to 1ug); max volume of RNA = 17.75uL. Generally, identify the RNA sample with the lowest concentration and multiply by 17.75uL. Use this quantity (ug) of RNA for each and every sample.
2. Transfer calculated volume(s) of RNA to 0.5mL snap cap tubes or PCR plate. Adjust volumes of individual samples to 17.75uL with H2O.
3. Add appropriate amount of primer to sample. Use 0.25ug primer per 1ug of RNA in sample (= 0.5uL of Promega oligo dT Cat#C1101 in this example). Total volume (RNA + primers) should equal 18.25uL.
4. Heat samples at 70C for 5 min in thermocycler.
5. Place samples on ice IMMEDIATELY.
6. Make Master Mix:
PER RXN
5 uL 5x Buffer (M-MLV RT Buffer)
1.25 uL 10mM dNTPs (Promega Cat#U1511)
0.5 uL M-MLV RT per ug of RNA
Used 4uL of RNA (
1ul of rna from each of the original sea pen sample added to a single sample combining rna sp1-1 sp1-2 sp2-1 sp2-2)
7. Mix well.
8. Add 6.75uL of master mix to each reaction.
9. Mix well, but do not vortex.
10.Spot spin.
11.Incubate @ 42C for 1hr in thermalcycler for oligo dT primers OR @ 37C for random primers.
12.Heat inactivate @ 95C for 3 min.
13.Spot spin.
14.Store @ -20C.
Sucessfully converted RNA to cDNA next step is to replicate using PCR and verify on a gel.
Subscribe to:
Posts (Atom)