Objective: Verify Sea Pen luciferase sequence was sucessfuly replicated using primerse based on Renilla Reniformes luciferase sequence acquired on Genebank.
PCR gel run today
Sea pen 1 and 2
1-1 lane 2 (note: Made sea pen luminescence before taking samples on both 1-1 (base cut) and 1-2 (top cut) )
1-2 lane 3 (note: Made sea pen luminescence before taking samples on both 1-1 1-2)
2-1 lane 4
2-2 lane 5
- lane 6
- lane 7
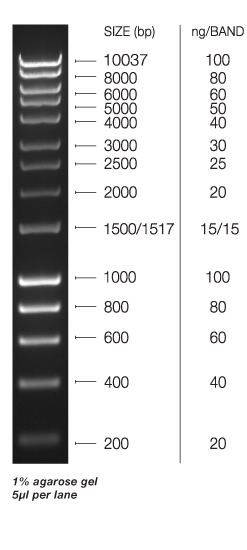
Bioline hyper ladder 1
Joined Dr. Roberts class lab and performed a conventional pcr (2x Apex Red)
6 tube 1-1,1-2,2-1,2-2,and 2 negative control tubes
master mix
2x apex red 12.5ul*6 =75ul
f primer 0.5ul*6 3ul
r primer 0.5ul*6 3ul
h20 6.5ul*6 39ul
=20ul per tube
template 5ul*6 30ul
95c-10 min
40 cycles of
95c 15s
50c 15s
72c 1min
Results: Replication was successful removed the brightest bands in lane 3 and 4 from the gel, and purified to prepare for sequencing. Samples were sent to "" (will add later)
Found out that the problem with the reverse transcription quantities was a pipetting error on the 10ul pipet i was using to much because of a mix up on the decimal place of the pipette
Introduced a spacer for the sea pens to keep them from touching each other testing it on the 5 gallon bucket . It splits the bucket in half reducing the area they are able to move around in without blocking flow or height the pen can extend Will see if it improves individual health of the sea pens.
With the automatic feeder I have noticed that the Pens have become much more consistent in activity so I believe I am set it up so that all of them can be sustained for a long period of time though more time will be needed to tell if this is true. It may be beneficial to create a paper just on my setup which could help others working with sea pens or related species.